Sourdough Helps Syracuse Scientists Understand Evolution
Editor’s Note: The following article was written by Cameron Murray, a research technician in biology professor Angela Oliverio’s lab.
No matter how alone we may feel at times, we are always kept company by microbial life. Microorganisms are ever present in our bodies, our homes, in the air we breathe, and the food we eat. These microbial interactions shape much more of our daily life than we realize.
With “microbiome” becoming a buzzword, there has been a strong push to study the microbes that shape our health and environment, yet understanding these microbes at a real-world, community-level remains out of grasp. In the lab, scientists tend to study microbes in precisely controlled, isolated cultures, providing insight into individual organisms outside of their natural environment. Conversely, scientists in the field cast a wide net in surveys of environmental microbes, capturing diversity but obscuring community functions.
Angela Oliverio, an assistant professor of biology in Syracuse University’s College of Arts and Sciences (A&S), proposes a new model system to bridge this gap: fermented foods. With a five-year, $2.1 million MIRA (Maximizing Investigators’ Research Award) grant from the National Institutes of Health, she will investigate how microbial communities evolve using sourdough starters.

Fermented foods like sourdough, kimchi and kombucha contain whole communities of interacting microbes but are still manageable enough to study in the lab. A fermented food microbiome may contain around three to 10 different species, while a single human gut sample contains hundreds. Fermented foods strike a balance between the simplicity of single-organism lab experiments and complex microbiome analyses.
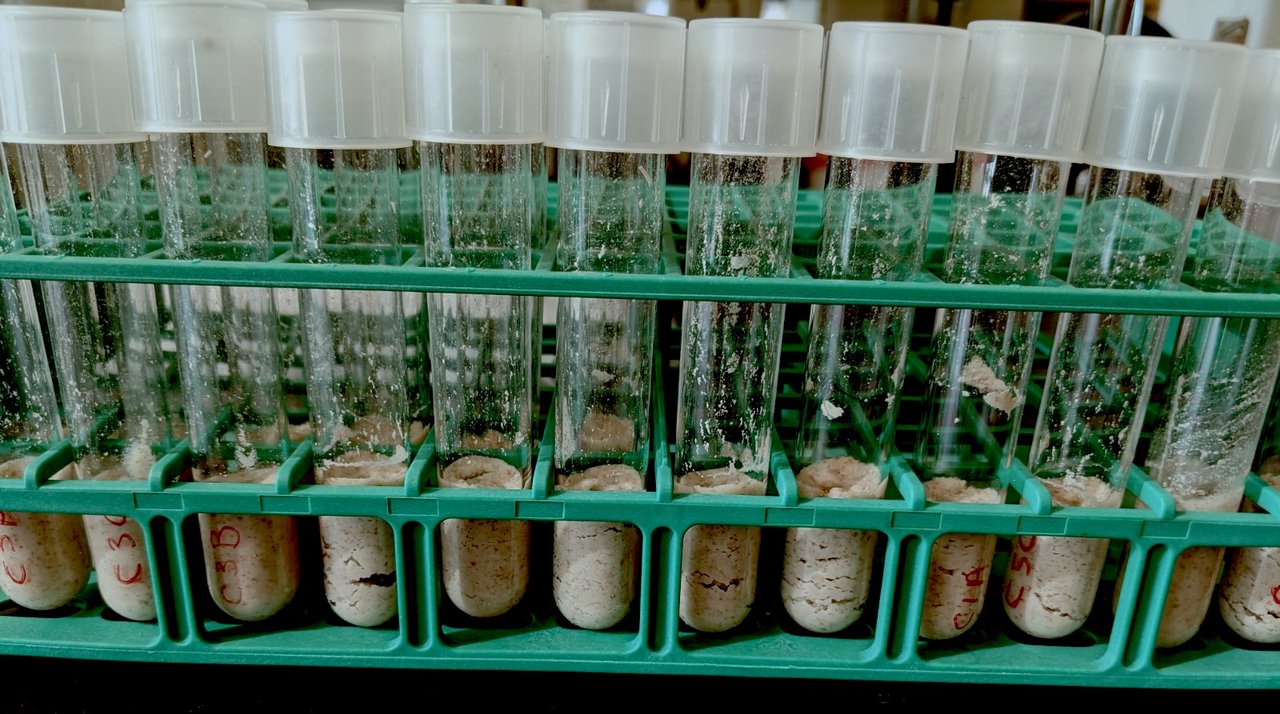
Sourdough in particular is well suited for this research because of the way a starter is maintained. These fermented cultures are repeatedly passaged from the previous batch into the next, or in baker’s terms, feeding the starter. All starters share functional groups of microbes, but each individual is unique in the diversity of species and strains present, providing an ideal environment for tracking how microbial communities change and evolve over time.
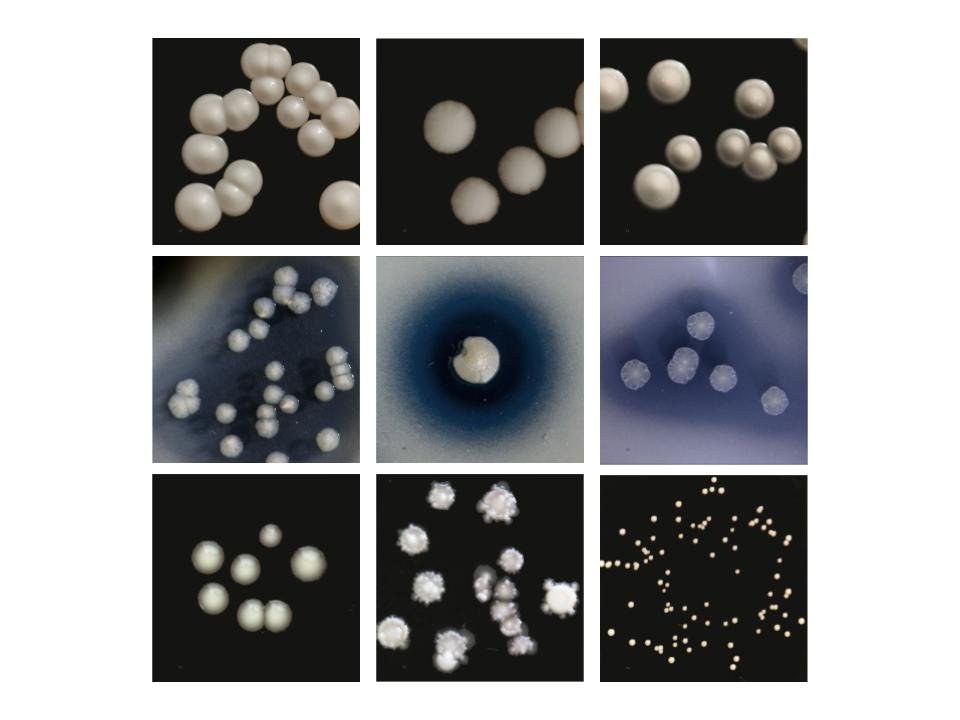
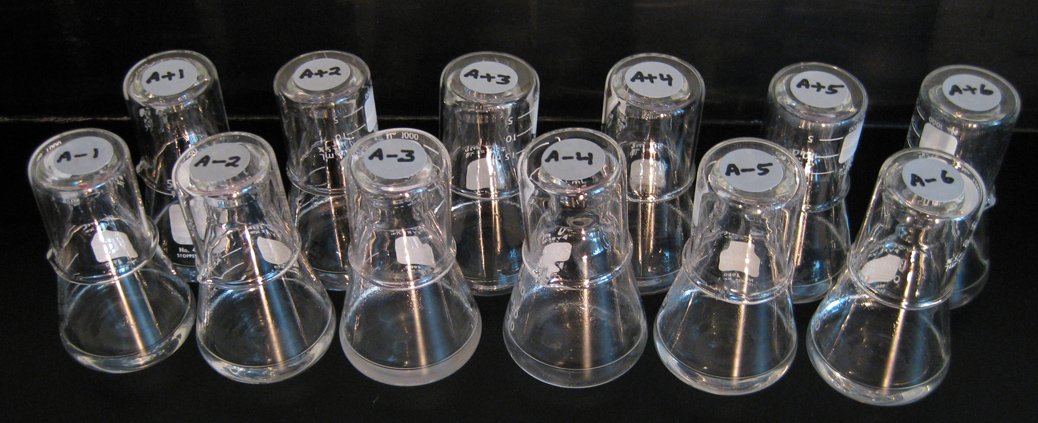
Oliverio’s work builds upon the legacy of one of biology’s most famed experiments, the E. coli Long-Term Evolution Experiment (LTEE), in which 12 initially identical strains of E. coli have been maintained since 1988, surpassing 80,000 generations in 2024. The E. coli LTEE revealed the mechanisms behind evolutionary change within a single species in isolation. Notably, one of the cultures evolved the ability to use citrate, a nutrient source the ancestral strain could not use. Oliverio intends to understand how these evolutionary processes work in a multispecies context.

Previous research in the Oliverio lab showed that small, strain-level differences among microbes can have a huge impact on microbial community composition and functional outputs in fermented foods like flavor and texture.
By constructing microbial communities in the lab that differ only in the strain of a single member, Oliverio’s team can track how those communities evolve over time. Oliverio plans to illuminate the evolutionary processes that lead to strain divergence in a realistic fermented food context. The team will then combine data from these experiments with an extensive collection of microbial genomes to build a computational model that can predict the outcomes of different synthetic microbial communities. This model would be useful for scientists in myriad contexts.

At a basic science level, Oliverio’s research will deepen our understanding of how evolution operates in the kinds of complex, multispecies environments that characterize nearly every ecosystem on Earth, including the human body. On a more applied level, insights from these experiments could inform improvements to fermented food production and help scientists design more effective probiotics.
"Sourdough is something people keep in their kitchens and feed, often for years or even decades," Oliverio says. "These microbial communities have been evolving alongside humans for thousands of years. That we can use them to understand cutting-edge questions in biology is pretty amazing. And people love to tell me about their starters."
Conflict of interest statement: The author of this article conducted some of the preliminary experiments described in the grant proposal.
Published: May 11, 2026
Media Contact: asnews@syr.edu
